|
The telomerase enzyme contains a catalytic part and a built-in RNA template. 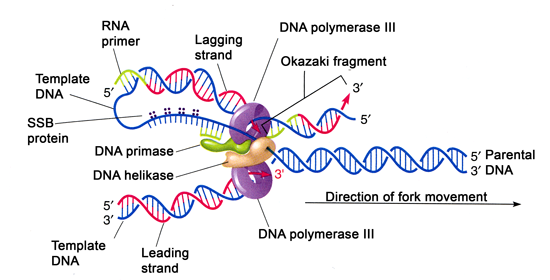
The telomeres are added to the ends of chromosomes by a separate enzyme, telomerase ( Figure), whose discovery helped in the understanding of how these repetitive chromosome ends are maintained. In a way, these telomeres protect the genes from getting deleted as cells continue to divide. In humans, a six-base-pair sequence, TTAGGG, is repeated 100 to 1000 times in the telomere regions. Telomeres comprise repetitive sequences that code for no particular gene. The DNA at the ends of the chromosome thus remains unpaired, and over time these ends, called telomeres, may get progressively shorter as cells continue to divide. When the replication fork reaches the end of the linear chromosome, there is no way to replace the primer on the 5â end of the lagging strand. On the lagging strand, DNA is synthesized in short stretches, each of which is initiated by a separate primer. In the leading strand, synthesis continues until the end of the chromosome is reached. As youâve learned, the enzyme DNA pol can add nucleotides only in the 5' to 3' direction. Unlike prokaryotic chromosomes, eukaryotic chromosomes are linear. The gaps that remain are sealed by DNA ligase, which forms the phosphodiester bond. The Okazaki fragments in the lagging strand are joined after the replacement of the RNA primers with DNA. The displaced primer RNA is then removed by RNase H (AKA flap endonuclease) and replaced with DNA nucleotides. 
As pol δ runs into the primer RNA on the lagging strand, it displaces it from the DNA template. A sliding clamp protein known as PCNA (proliferating cell nuclear antigen) holds the DNA pol in place so that it does not slide off the DNA. While the leading strand is continuously synthesized by the enzyme pol δ, the lagging strand is synthesized by pol ε. DNA pol α adds a short (20 to 30 nucleotides) DNA fragment to the RNA primer on both strands, and then hands off to a second polymerase. Three major DNA polymerases are then involved: α, δ and ε. Primers are formed by the enzyme primase, and using the primer, DNA pol can start synthesis. These are resolved with the action of topoisomerases. The opening of the double helix causes over-winding, or supercoiling, in the DNA ahead of the replication fork. Replication forks are formed at each replication origin as the DNA unwinds. Difference between Prokaryotic and Eukaryotic ReplicationĪ helicase using the energy from ATP hydrolysis opens up the DNA helix. Helicase and other proteins are then recruited to start the replication process ( Table). At the origin of replication, a pre-replication complex is made with other initiator proteins. The chromatin (the complex between DNA and proteins) may undergo some chemical modifications, so that the DNA may be able to slide off the proteins or be accessible to the enzymes of the DNA replication machinery. Histones must be removed and then replaced during the replication process, which helps to account for the lower replication rate in eukaryotes. Eukaryotic DNA is bound to basic proteins known as histones to form structures called nucleosomes. Before replication can start, the DNA has to be made available as a template. The essential steps of replication are the same as in prokaryotes. They are known as pol α, pol β, pol γ, pol δ, and pol ε. The number of DNA polymerases in eukaryotes is much more than in prokaryotes: 14 are known, of which five are known to have major roles during replication and have been well studied. 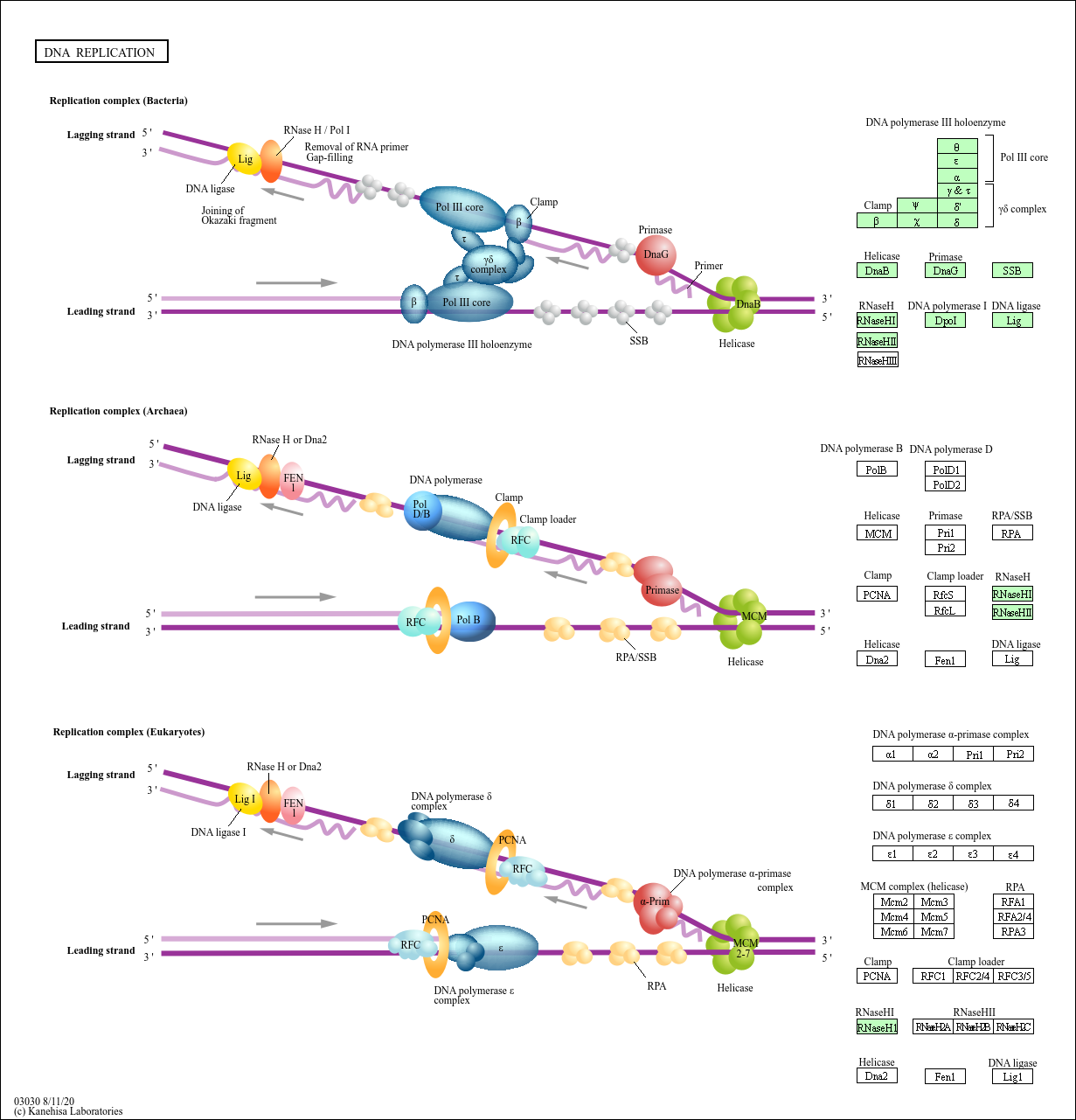
These are equivalent to the origin of replication in E. In yeast, which is a eukaryote, special sequences known as autonomously replicating sequences (ARS) are found on the chromosomes. The rate of replication is approximately 100 nucleotides per second, much slower than prokaryotic replication. 
There are multiple origins of replication on each eukaryotic chromosome humans can have up to 100,000 origins of replication across the genome. The human genome has 3 billion base pairs per haploid set of chromosomes, and 6 billion base pairs are replicated during the S phase of the cell cycle. Eukaryotes also have a number of different linear chromosomes. Eukaryotic genomes are much more complex and larger in size than prokaryotic genomes.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
